Add your promotional text...
The Epigenetic Bridge: How Embryos Inherit the Blueprint for Life
Meghana S. Oak, Marco Stock, Ana Janeva, Matthias Mezes, Antony M. Hynes-Allen, Tobias Straub, Ignasi Forné, Andreas Ettinger, Stephan Hamperl, Axel Imhof, Jelle van den Ameele, Antonio Scialdone & Eva Hörmanseder
RESEARCH ARTICLE
1/4/20262 min read
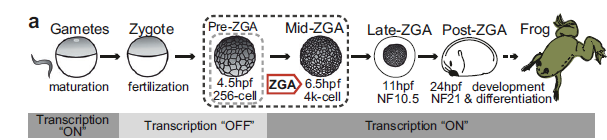
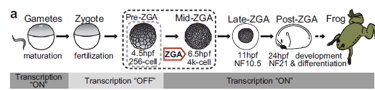
For decades, the prevailing narrative of early development was that the zygote began as a "blank slate." It was believed the genome sat idle until maternal transcription factors arrived to jumpstart the engine of life during Zygotic Genome Activation (ZGA). This study effectively upends that model, proving that embryos don't actually start from zero. Instead, they rely on a remarkably stable "pre-marking" system—specifically the histone mark H3K4me3, which is established in the gametes and preserved through the silent phases of early cleavage.
This discovery reveals that the maternal-to-zygotic transition isn't a sudden hand-off of control, but a continuous epigenetic bridge. By demonstrating that H3K4 methylation is a prerequisite for transcription rather than just a byproduct of it, the researchers have pinpointed a core mechanism of developmental "memory." It suggests that the embryo's ability to execute its complex program is "primed" long before the first gene is ever turned on.
Ultimately, this shifts our perspective on cell potency and developmental failure. If the inherited epigenetic map is "blurred" or missing these H3K4me3 marks, the embryo cannot find its way, regardless of which transcription factors are present. This achievement places epigenetic inheritance at the very center of the logic that allows a single cell to faithfully build a complex organism.
To track these epigenetic shifts, the team used Xenopus laevis (African clawed frog), leveraging its large embryos and clear ZGA timeline. They employed a multi-omic strategy to correlate chromatin structure with gene activity:ChIP-seq and CUT&RUN: Mapped H3K4me3 and other histone marks across gametes and embryos both before and after ZGA.
RNA-seq: Used to verify exactly when gene expression began in relation to those marks.
Loss-of-Function Assays: Targeted the "writers" of these marks (Cxxc1 and Kmt2b) using Morpholino-mediated knockdown to see if the embryo could function without them.
Timed Demethylation: Stripped H3K4me3 at specific intervals to determine exactly when this "marking" is most vital for survival.
Scientific Breakthroughs & Key Results
Inheritance Without Activity: The study identified thousands of genomic sites where H3K4me3 was present in gametes and maintained in pre-ZGA embryos, despite a complete absence of transcription. These marks were found predominantly in CpG-rich, hypomethylated regions.
Essential for Master Regulators: When Cxxc1 and Kmt2b were knocked down, H3K4me3 levels plummeted at the promoters of critical pioneer factors. Most notably, the expression of Pou5f3.2 (the Oct4 homolog) and Sox3 failed, leading to a total collapse of gastrulation.
The Critical Window: Experiments revealed that removing H3K4me3 before ZGA (during early cleavage) was just as lethal as removing it during active transcription. This confirms that the pre-marking phase is a non-negotiable bottleneck for development.
Breadth as a Signature: The researchers found that these inherited "memory" sites are distinguished by their physical breadth, often spanning several kilobases. This "broad domain" signature acts as a clear marker for genes that must be activated first.
While this study identifies H3K4me3 as the primary architect of ZGA, it opens new questions about how these marks are placed. We still don't fully understand how Cxxc1 and Kmt2b are recruited to specific CpG sites when the genome is silent, is it written in the DNA sequence itself, or are there undiscovered "pioneer" proteins guiding the way?
Furthermore, the discovery that Xenopus patterns mirror human oocytes more closely than mouse models suggests we need a broader comparative look across species to see if this "bridge" is a universal rule for all vertebrates. Finally, there is the question of "epigenetic erosion": could environmental stressors or parental age degrade these marks, leading to developmental pathologies long before a heart even begins to beat?